There is intriguing evidence for some ancient genes giving rise to very different eye types, including our own eyes as much as those of invertebrates. The best-known example for that is the gene Pax-6, which is an essential early eye development gene in both mice and flies. The genes are so similar in these evolutionarily extremely distant animals that deficient flies even can be rescued by the mouse gene, and vice versa. Many more orthologous genes exist in mice and flies, and there are many research labs that study how these genes function in either mice or flies. More generally, we know a lot about how the eyes of a few species, such as fruit flies and mice develop, but we know very little to what extent such developmental patterns can be generalized. Among arthropods eye development research tends to focus on the fruit fly Drosophila melanogaster, yet there are around 1 million species of different arthropods (~90% of which are insects) inhabiting vastly different visual-ecological niches. The question of how generalizable known eye development patterns are, and how genetic pathways may be altered to accommodate functional specializations is a very exciting one.
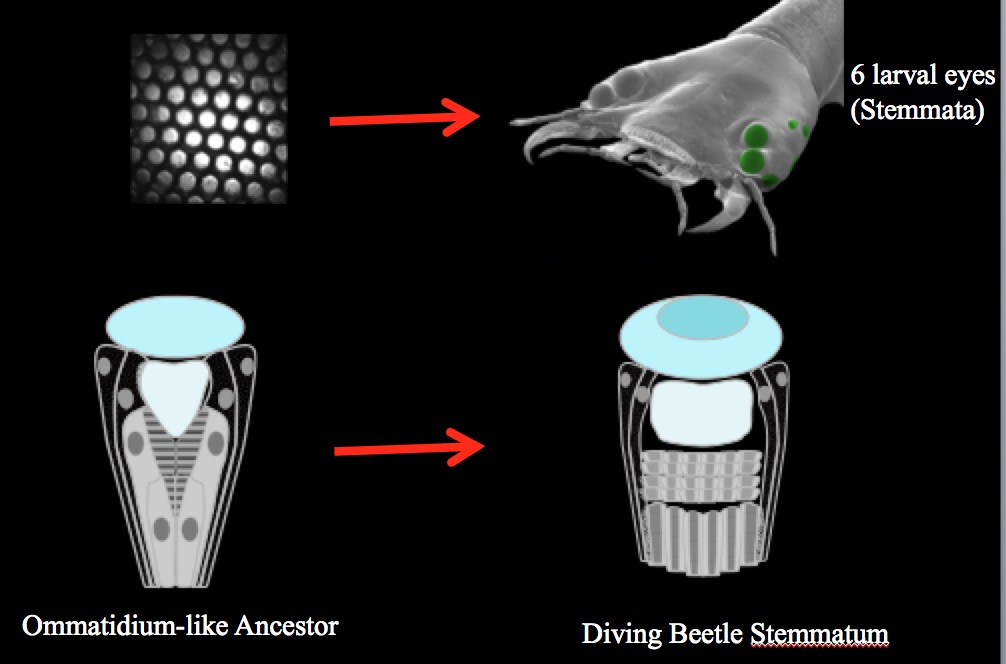
To address this fundamental question we investigate which, where and when known key eye development genes are expressed in the image-forming eyes of diving beetle larvae. These eyes are particularly interested because they are functionally sophisticated and they likely evolved from an ancestral compound eye.
There are many reasons why we believe that the bizarre beetle larval eye evolved from a compound-eye like ancestor. For a detailed explanation, please see Buschbeck (2014). Consistent with that notion we found that similar to components of the Drosophila adult compound eye, the beetle larval eyes (stemmata) all develop from a progenitor epithelium by recruiting comparable cell types in a comparably sequential manner. Based on these morphological similarities we established a cell-type homology model that allows us to make very specific predictions in regards to when and where specific genes might be active. A large portion of this project is to test these predictions.
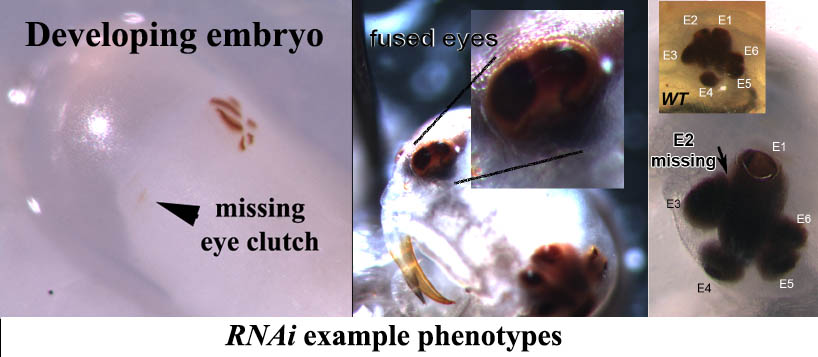
We also use RNAi, a technique that allows us to knock down the expression of specific genes and to observe if and how resulting gene deficits alter the morphological development of the diving beetle larval eyes. Here too we can test very specific expectations that are based on our knowledge on how specific genes contribute to Drosophila eye development. So far our gene knock-downs have lead to many different phenotypes, including animals that are missing the entire eye clutch, animals in which several eyes fuse into a single unit, and animals that miss individual eyes.
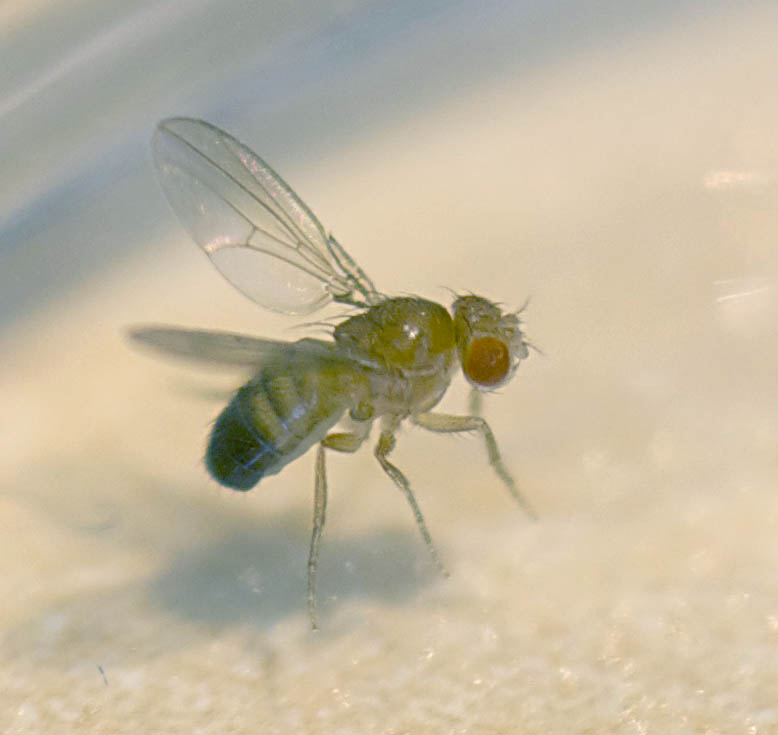

To further explore the function of specific genes we also take advantage of the genetically powerful fruit-fly model. Here it is possible to test the effect of a specific mutation in a cell specific manner. Our collaborator, Dr. Tiffany Cook at the Center for Molecular Medicine and Genetics at Wayne State University is studying the relationship between a particular glia-like eye cell and photoreceptors. In her lab she engineers flies that have very specific deficiencies and we then test them in regards to their physiology and optics.
Lab members primarily associated with these projects: Aaron Stahl (recently graduated), Arun Muthusamy, Winter Partin, Taryn Miller, Mike Le, Avery Wisenbarger.

Large parts of this project are funded by the National Science Foundation under Grant IOS- 1456757 to Dr. Elke Bushbeck with Co-PI Dr. Tiffany Cook